Please check back soon for more information as we are constantly updating our file descriptions based on search frequency.
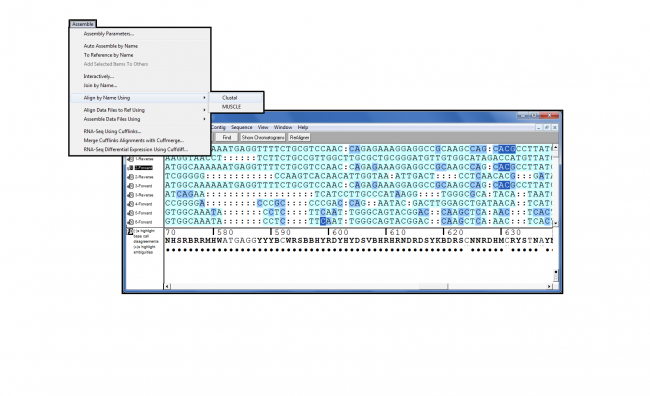
Extensive data import & export capabilities, inc. Educational institutions use Sequencher or other similar program, but they can be so expensive they. that are vital to performing research such as printing, saving, and exporting analyses. The transcripts.txt file contains the list transcripts IDs that I want to export (both the IDs and the sequences) from assembly.fasta to selectedtranscripts.fasta. Sequencher is the biologists choice for DNA sequence assembly and analysis. most popular one is the Standard Chromatogram File, or scf. We have yet to investigate this file type further, or there was not enough information available at the time to report accurately on the format. I want to extract specific fasta sequences from a big fasta file using the following script, but the output is empty. This data file format was added to our database by a visitor to this site, but no additional information was provided. from Bio import SeqIO records SeqIO.parse ('THISISYOURINPUTFILE.fasta', 'fasta') count SeqIO.write (records, 'THISISYOUROUTPUTFILE.nexus', 'nexus') print ('Converted i records' count) Or you can use this site as online.
#SEQUENCHER EXPORT FASTA AS ONE FILE HOW TO#
If you are unable to open the file this way, it may be because you do not have the correct application associated with the extension to view or edit the FASTA file. How to convert from fasta to nexus You can also convert between these formats by using command line tools.

Open the file and copy paste the sequences into BOLD. You should now have a text file in your folder with. In the Export Subproject box, beside Export Location press Browse to find your file. The best way to open an FASTA data file is to simply double-click it and let the default assoisated application open the file. Go to File > Export> Selection As Subproject. If you are aware of any additional file formats that use the FASTA extension, please let us know. Click on the link to get more information about listed programs for export fasta file action. Programs supporting the exension fasta on the main platforms Windows, Mac, Linux or mobile. FASTA extension are known as FASTA Sequence files, however other file types may also use this extension. Applications that export fasta file - FASTA format DNA and protein sequence alignment.
#SEQUENCHER EXPORT FASTA AS ONE FILE SOFTWARE#
Have you found, downloaded or received an FASTA file, but don't know which software program is required to open it?īefore attempting to open an FASTA file, you'll need to determine what kind of file you are dealing with and whether it is even possible to open or view the file format.Īnswer: Files which are given the.
